Even though matrices have been used extensively for dynamic programming, adjacency matrix representation of dense graphs and many other essential use-cases, ML workloads live and breathe matrices.
Since usually dimensionality is not known at compile time, dynamic arrays are usually used as a primitive. As such I often see something like
to represent NxM matrix. While it may seem natural and intuitive, such representation comes with a number of shortcomings. The most obvious issue with this is the waste of memory to store all 3 member fields of N nested vectors. The other issue is not obvious but potentially more impactful in terms of performance - each nested vector has its own chunk of heap memory that may be located on random virtual memory pages and have to be accessed through pointer indirection.
Fortunately there is really no reason to use such nested vector setup and a one-dimensional vector is more than enough:
Now we just need to perform index computation ourselves instead of letting compilers do it, so instead of something like
we have
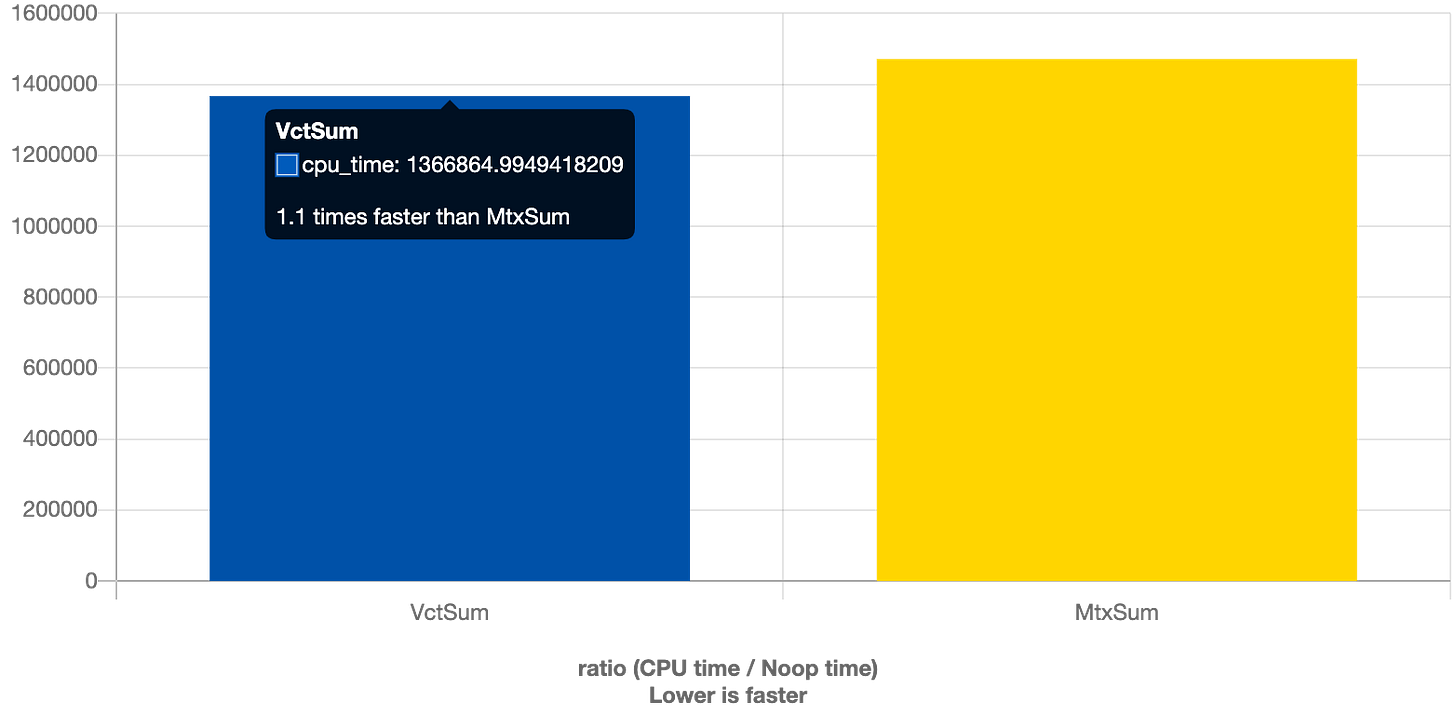
Does it matter though?
Looks like there is a 10% difference for 100000 X 10 matrix but it grows with the number of nested vectors, so it may be somewhat mitigated by swapping dimensions if number of rows is much larger than the number of columns.
Since working with more dimensions becomes even more cumbersome, most ML frameworks have special primitives that provide multi-dimensional views on top of single-dimensional vectors and C++23 finally makes this very easy as well with std::mdspan.
So next time you have to work with tensors, check your libraries if they provide mdspan-like abstractions and if not, consider writing your own.